Genomics research shows that more than a million proteins are encoded by approximately 30,000 human genes. Proteomics, the study of proteins encoded by the genome, includes identification of post-translational modifications, structural analyses, protein localization studies, and protein quantitation. Mass spectroscopy-based techniques have evolved as a powerful tool in proteomics. Stable isotope-labeled peptides (SIL peptides) are chemically and physically indistinguishable from their endogenous counterparts with respect to retention time, ionization efficiency, and fragmentation pathways. Therefore, either are ideal internal standards and template analytes. Peptides can be labeled with one or more isotopes of hydrogen, carbon, nitrogen, or oxygen by incorporating amino acids containing the desired isotopes such as deuterium (D), 13C, 15N, or 18O into the peptide during synthesis.
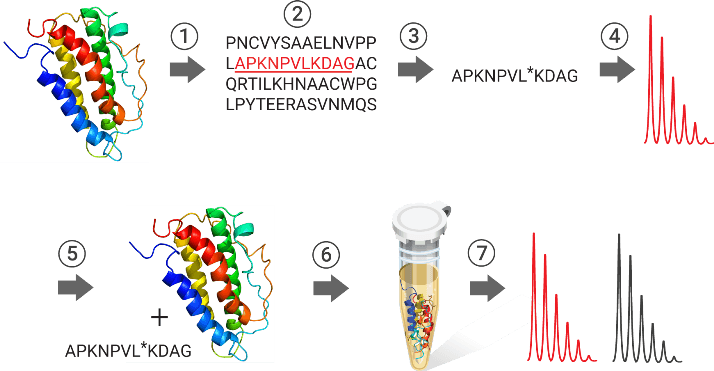
The Absolute Quantification method (AQUA) enables targeted quantification of protein and post-translational modifications in complex protein mixtures using SIL peptides as internal standards. The SIL peptide is introduced into a biological sample during or after protease digestion. The heavy SIL peptide and its light endogenous peptide fragment are detected by selected reaction monitoring (SRM) in a mass spectrometer. Based on the known amount of SIL peptide added, and the intensity ratio of both peptides, the amount of endogenous protein can be calculated.
Isotopic Labeling Chemistry
Isotopic Amino Acid Table
| AMINO ACID | ISOTOPE | MASS DIFFERENCE | ISOTOPIC ENRICHMENT |
|---|---|---|---|
| Alanine | 13C3, 15N | 4 | >99% |
| Arginine | 13C6, 15N4 | 10 | >99% |
| Isoleucine | 13C6, 15N | 7 | >99% |
| Leucine | 13C6, 15N | 7 | >99% |
| Lysine | 13C6, 15N2 | 8 | >99% |
| Phenylalanine | 13C9, 15N | 10 | >99% |
| Proline | 13C5, 15N | 6 | >99% |
| Valine | 13C5, 15N | 6 | >99% |
SIL Peptides in Structural Analysis
Nuclear Magnetic Resonance (NMR) is a powerful technique for investigating structural information, dynamics, and molecular interactions of biomolecules. This technique can be used to measure relaxation rates of biomolecules as they dissociate from their bound target. Peptides labeled with D (spin of 1), 13C (spin 1/2), and 15N (spin 1/2) are suitable for NMR studies of proteins. A combination of NMR spectroscopy and segmental isotopic labeling is used to study the mechanism of protein splicing (e.g., the structure of an active protein splicing precursor), a post-translational autocatalytic process in which an intervening sequence, termed an intein, is removed from a host protein, the extein.
High Quality SIL Peptides from CPC Scientific
Absolute quantitation of a complex protein mixture at very low concentrations and structural studies requires high-quality peptides enriched with stable isotopes. The CPC Scientific SIL Peptide Custom Synthesis Service guarantees superior quality and high isotopic enrichment. These stable isotopic peptides are synthesized using the latest Fmoc solid-phase peptide technology in our state-of-the-art peptide laboratory. All heavy isotope-labeled peptides undergo mass spectrometric analysis and stringent analytical HPLC to establish the final purity and assure that our customers receive only the highest quality peptides for absolute quantitation and other studies.
Two major isotopes, 15N and 13C, are incorporated into a specific amino acid sequence of peptides. Each atom contains over 99% of an enriched isotope and can be located at multiple positions within the peptide. For example, a Leu amino acid containing one 15N and six 13C, and a peptide containing one such isotope-labeled Leu amino acid, has seven units of molecular weight higher than the corresponding unlabeled peptide.
SIL Peptide Applications
- Functional quantitative proteomics
- Quantitation of post translationally-modified proteins
- Protein structure analysis
- Protein expression monitoring
- Biomarker discovery
- Pharmacokinetics
- Metabolomics
- Clinical biochemistry for drug and metabolite monitoring
- Anti-doping testing
- Cell signal profiling and pathway validation
- Protein cross-linking analysis
SIL Peptide Modifications
- Phospho-Tyr, Ser, Thr (single or multiple)
- Sulfo-Tyr (single or multiple)
- Methylated Arg, Lys
- Chloro-Tyr
- Met-oxidized
- Pyroglutamic acid
The Absolute Quantification method (AQUA) enables targeted quantification of protein and post-translational modifications in complex protein mixtures using SIL peptides as internal standards. The SIL peptide is introduced into a biological sample during or after protease digestion. The heavy SIL peptide and its light endogenous peptide fragment are detected by selected reaction monitoring (SRM) in a mass spectrometer. Based on the known amount of SIL peptide added, and the intensity ratio of both peptides, the amount of endogenous protein can be calculated.
Isotope-Labeled Peptide Citations
Development of a selective, sensitive and robust oxyntomodulin dual monoclonal antibody immunoassay.
Umberger, T.S., Ming, W., Cox, J.M., Konrad, R.J. and Siegel, R.W. Bioanalysis 14, no. 18 (2022): 1229-1239.
- Lilly Research Laboratories, Eli Lilly & Company, Indianapolis, IN46285, USA
Human K2 EDTA and P800 plasma (500 μl) was spiked with proglucagon 33–61, 35–61 and 36–61 stable-isotope-labeled internal standard peptides (CPC Scientific, custom order) and diluted with I buffer (25 mmol/l Tris-HCl, 25 mmol/l HEPES, 300 mmol/l NaCl, 0.1% (v/v) octyl β-D-glucopyranoside, pH 7.5).
Coskun, T., Urva, S., Roell, W.C., Qu, H., Loghin, C., Moyers, J.S., O’Farrell, L.S., Briere, D.A., Sloop, K.W., Thomas, M.K. and Pirro, V. Cell Metabolism 34, no. 9 (2022): 1234-1247.
- Eli Lilly and Company, Lilly Corporate Center, Indianapolis, IN 46285, USA
Homologous and heterologous competition experiments were performed with non-radioactive peptide analogues[127I]-Tyr1-GIP(1-42) and [127]-Tyr10-GIP(1-42) to ensure quantification of the high-affinity binding site of the GIPR. Peptide analogues were generated using synthetic [127I]-Tyr amino acid building blocks (CPC Scientific).
Libiger, O., Shaw, L.M., Watson, M.H., Nairn, A.C., Umaña, K.L., Biarnes, M.C., Canet‐Avilés, R.M., Jack Jr, C.R., Breton, Y.A., Cortes, L. and Chelsky, D. Alzheimer's & Dementia 17, no. 12 (2021): 1976-1987.
- Janssen Research and Development, 3210 Merryfield Row, San Diego, CA 92121, USA.
- Foundation for the National Institutes of Health, North Bethesda, Maryland, USA
- Department of Radiology, Mayo Clinic, Rochester, Minnesota, USA
- Merck & Co., Inc., West Point, Pennsylvania, USA
Purified synthetic stable isotope (15N and 13C) labeled (SIL) peptides were from CPC Scientific (EDSLEAGLPLQVR [CMGA]; SLGVGFATR [FABPH]; VLEYLNQEK [SCG2]; and THLGEALAPLSK [VGF]); and JPT Peptide Technologies (TNYLYGK [NPTX2]).
Pantua, H., Skippington, E., Braun, M.G., Noland, C.L., Diao, J., Peng, Y., Gloor, S.L., Yan, D., Kang, J., Katakam, A.K. and Reeder, J. MBio 11, no. 5 (2020): 10-1128.
- Departments of Infectious Diseases, OMNI Bioinformatics, Chemistry, Structural Biology, Biochemical and Cellular Pharmacology, Translational Immunology, Pathology, Molecular Biology, Genentech, South San Francisco, California, USA
MQLNKV-L(U13C6,15N)-KGL(U13C6,15N)MIALPVMAIAA-dipalmitoyl2C-SSNKNGG-K-biotin, which upon cleavage by LspA, yields the product peptide dipalmitoyl2C-SSNKNGG-K-biotin. A product standard [dipalmitoyl2C-SSNKNAAK-(NHCH2CH2NH)-biotin; CPC Scientific] was in the reaction mixture as an internal standard for normalization of product quantitation.
Bårdsen, Kjetil, Michaela D. Gjerstad, Markku Partinen, Ingeborg Kvivik, Anne Bolette Tjensvoll, Peter Ruoff, Roald Omdal, and Cato Brede. Analytical Chemistry 91, no. 14 (2019): 9323-9329.
Synthetic Hcrt1 with13Cand15N stabile isotopemodification on two leucine amino acids was used for internalstandard (ISTD) calibration; Glp-P-L(U13C6,15N)PDCCR-QKTCSCR-L(U13C6,15N)YELLHGAGNHAAGILTL-NH2(CPC Scientific, Sunnyvale, CA)
Dekker, Petrus Jacobus Theodorus, René Marcel De Jong, and Cornelis Marinus Muijlwijk. U.S. Patent Application No. 14/397,866.
"The internal standard (ISTD) WIL*GDVFIREYYSV*FDR was ordered from CPC Scientific (Sunnyvale, Calif., USA) and contained L*6×13C 1×15N and V*5×13C 1×15N. The ISTD contained an internal trypsin cleavage site in order to monitor the digestion with trypsin to completion."
Cox, J.M., Berna, M.J., Jin, Z., Cox, A.L., Sloop, K.W., Gutierrez, J.A. and Ackermann, B.L. Bioanalysis 8, no. 15 (2016): 1579-1595.
- 1 Lilly Research Laboratories, Eli Lilly & Company, Lilly Corporate Center, Indianapolis, IN 46285, USA.
"The OXM peptides 1–37, 3–37 and 4–37 as well as their corresponding stable isotope-labeled internal standards (each containing one [ 13 C 9 , 15 N]-labeled phenylalanine and two [ 13 C 6 , 15 N]-labeled leucine residues) were synthesized by CPC Scientific, Inc."
Zhen, Eugene Y., et al. Biochemical Journal (2015): BJ20151085.
- Lilly Research Laboratories, Lilly Corporate Center, Indianapolis, IN 46285, U.S.A
"Stable isotope-labelled (SIL) peptides were synthesized by CPC Scientific. Leucine residues were selectively labeled using 13C/15N-labelled amino acids resulting in the addition of 7 Da to the peptide mass (Supplementary Table S1). The reported purity of each synthesized peptide was more than 95% based on HPLC analysis. "
Lopez, A., Reyna, D.E., Gitego, N. et al. Co-targeting of BAX and BCL-XL proteins broadly overcomes resistance to apoptosis in cancer. Nat Commun 13, 1199 (2022).
- Cardio-metabolic Disease Biomarkers, Merck Research Laboratories, Kenilworth, NJ., Merck & Co., Inc.
"Glucagon synthetic peptide and heavy isotope–labeled internal standards (ISs) of oxyntomodulin and glucagon containing 2 [ 13 C 6 - 15 N]-labeled leucines and 1 [ 13 C 9 - 15 N]-labeled phenylalanine were synthesized by CPC Scientific.."
Adam, G.C., Meng, J., Rizzo, J.M., Amoss, A., Lusen, J.W., Patel, A., Riley, D., Hunt, R., Zuck, P., Johnson, E.N. and Uebele, V.N. Journal of Biomolecular Screening 20, no. 2 (2015): 212-222.
- Screening and Protein Sciences, Merck Research Labs, North Wales, PA, USA
Plates were incubated for 210 min at 25 °C followed by quenching with 25 uL water with 2% formic acid containing 600 nM of the internal standard with the sequence KTEEISEVN[L-U7]-OH (CPC Scientific).
